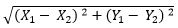K-Means Clustering Tutorial
During data analysis many a times we want to group similar looking or behaving data points together. For example, it can be important for a marketing campaign organizer to identify different groups of customers and their characteristics so that he can roll out different marketing campaigns customized to those groups or it can be important for an educational institute to identify the groups of students so that they can launch the teaching plans accordingly. Classification and clustering are two fundamental tasks which are there in data mining for long, Classification is used in supervised learning (Where we have a dependent variable) while clustering is used in un-supervised learning where we don’t have any knowledge about dependent variable.
Clustering helps to group similar data points together while these groups are significantly different from each other.
K-Means Clustering
There are multiple ways to cluster the data but K-Means algorithm is the most used algorithm. Which tries to improve the inter group similarity while keeping the groups as far as possible from each other.
Basically K-Means runs on distance calculations, which again uses “Euclidean Distance” for this purpose. Euclidean distance calculates the distance between two given points using the following formula:
Euclidean Distance =
Above formula captures the distance in 2-Dimensional space but the same is applicable in multi-dimensional space as well with increase in number of terms getting added. “K” in K-Means represents the number of clusters in which we want our data to divide into. The basic restriction for K-Means algorithm is that your data should be continuous in nature. It won’t work if data is categorical in nature.
Data Preparation
As discussed, K-Means and most of the other clustering techniques work on the concept of distances. They calculate distance from a specific given points and try to reduce it. The problem occurs when different variables have different units, e.g., we want to segment population of India but weight is given in KGs but height is given in CMs. One can understand that the distance matric discussed above is highly susceptible to the units of variables. Hence, it is advisable to standardize your data before moving towards clustering exercise.
Algorithm
K-Means is an iterative process of clustering; which keeps iterating until it reaches the best solution or clusters in our problem space. Following pseudo example talks about the basic steps in K-Means clustering which is generally used to cluster our data
- Start with number of clusters we want e.g., 3 in this case. K-Means algorithm start the process with random centers in data, and then tries to attach the nearest points to these centers
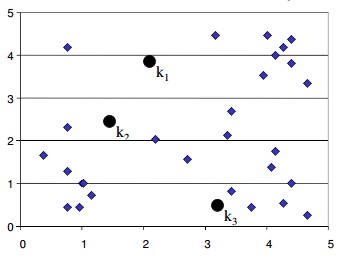
- Algorithm then moves the randomly allocated centers to the means of created groups
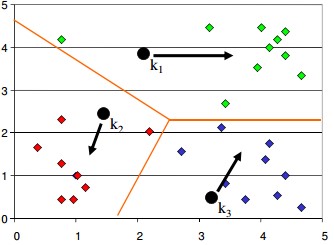
- In the next step, data points are again reassigned to these newly created centers
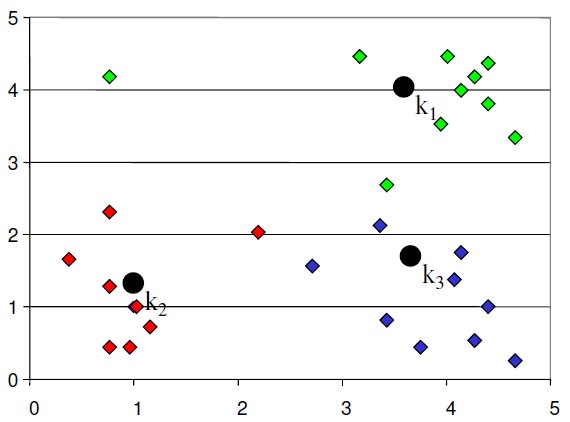
- Steps 2 & 3 are repeated until no member changes their association/ groups
Implementation in R
The step-wise pseudo code above explains about the K-Means clustering algorithm, but the basic question which should arrive is that how to decide the number of clusters (K)? How did someone identified that we need 3 clusters in above example? The objective of clustering exercise is to get the groups in a fashion that they are homogeneous within clusters and distinct from other groups. The very objective works as the solution to decide number of clusters “K” which we want to have in data. There is no mathematical formula which can directly give us answer to “K” but it is an iterative process where we need to run multiple iterations with various values of “K” and choose the one which is best fulfilling our purpose (As explained above).
Let’s use an example to understand this concept. We will be using “iris” dataset available in R to understand the same.
Get Closer To Your Dream of Becoming a Data Scientist with 70+ Solved End-to-End ML Projects
Iris dataset has following structure:

Now, let’s remove the last column from the data and keep only the numeric columns for the analysis
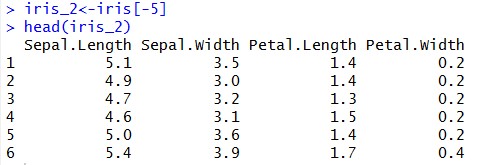
Once, we have dataset ready let’s standardize our data
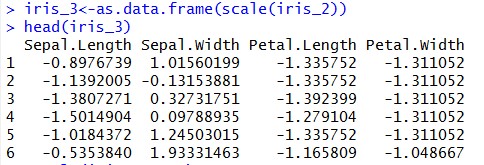
What have we achieved through scaling our data?
As we mentioned earlier that before executing K-Means on our data we have to standardize our data. Let’s check the ‘Mean’ & ‘Standard Deviation’ of “iris_2” & “iris_3” datasets:
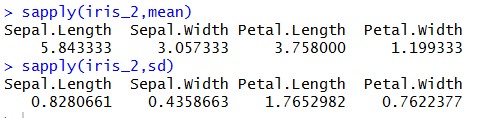
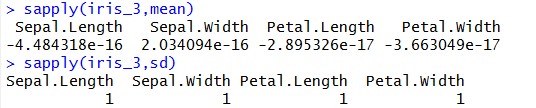
Looking at above results, you can understand that we have brought our variables to “Mean Zero – Standard Deviation – One”. During our implementation, we need to calculate “With-In-Sum-Of-Squares (WSS)” iteratively. WSS is a measure to explain the homogeneity within a cluster. Let’s create a function to plot WSS against the number of clusters, so that we can call it iteratively whenever required (Function name – “wssplot”, code is given at the end of this tutorial). We will be using “NbClust” library available at CRAN for this illustration.
Let’s first load the library-
> library(NbClust)
Deciding number of clusters “K”
Now, let’s call the “wssplot” function which we created.
> wssplot(iris_3, nc=30, seed=1234)
Here,
Iris_3 – dataset which we want to segment
nc – maximum number of clusters we are giving
seed – random initialization of clusters (Keep it constant if you want to retrieve the same clusters later point of time)
Output of above function is a plot which looks like:
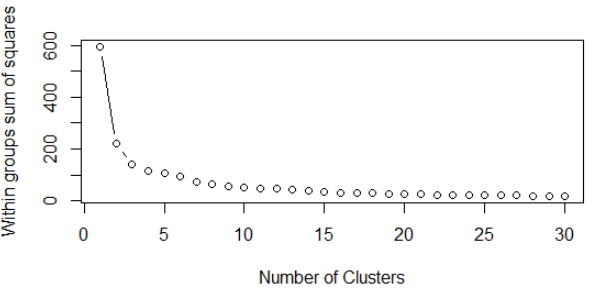
Here, we have plotted WSS with number of clusters. From here we can see that there is not much decrease in WSS even if we increase the number of clusters beyond 7. This graph is also known as “Elbow Curve” where the bending point (E.g, nc = 7 in our case) is known as “Elbow Point”. From the above plot we can conclude that if we keep number of clusters = 3, we should be able to get good clusters with good homogeneity within themselves. Let’s fix the cluster size to “7” and call the kmeans() function to give the clusters.
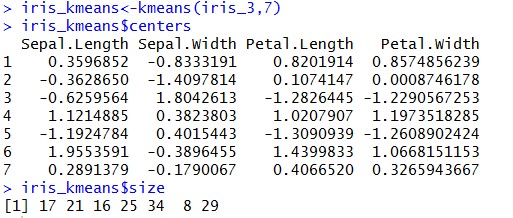
Above snippet shows that there are three clusters created with the centers as mentioned.
Get FREE Access to Machine Learning Example Codes for Data Cleaning, Data Munging, and Data Visualization
Let’s verify with original Species
We removed a last column from our iris dataset which had the specie type for each of the data-point. Let’s try to see how that classification is resonating with our clustering:
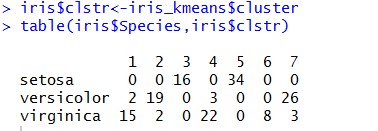
Seeing the results it seems that the clusters are quite good as our clusters don’t have mix of different species (except few exceptions).
Learn Data Science by working on interesting Data Science Projects
One more example using image reduction
I hope that you have understood a few concepts of K-Means clustering algorithm till now. As a reward it’s time do something interesting. Consider the image of a cute puppy below, what do you think how many colors are there in this image?
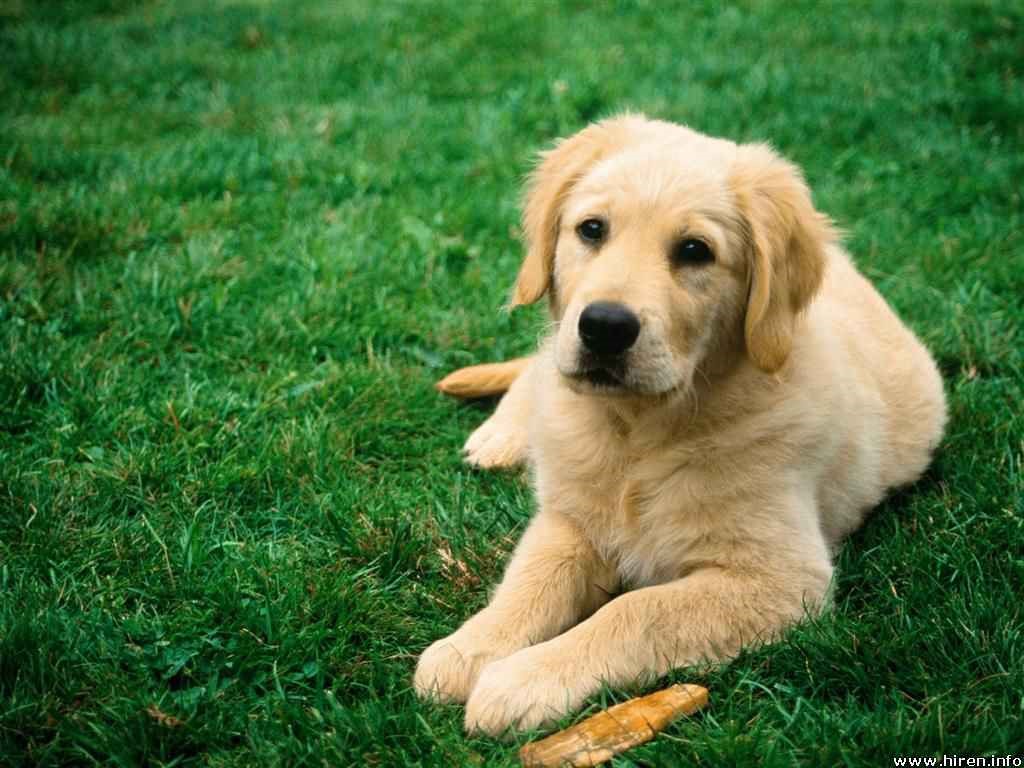

(Image source: Hiren.info)
This single image contains more than 124 K unique color combinations, our objective is to reproduce the similar image using very few colors (may be 3 or 4). Consider this image as a collection of three variables (R, G, B) at each pixel. To start, we need to load this image into R Session. We will be using “jpeg” package available in R to do this job for us.
Let’s get our image into a data frame format, i.e. we will try to get “Red”, “Green”, “Blue” value for each of the pixels
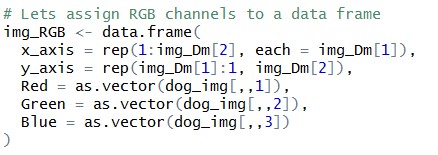
Img_RGB object has a structure which has R, G, B values at each and every pixel of our image. That means that now, image is nothing but a collection of three variables at each and every pixel.
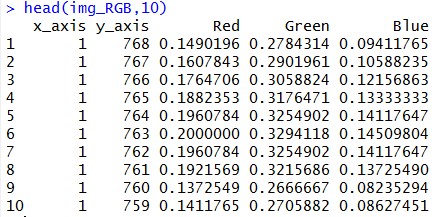
Let’s plot original image again using ggplot() function available in R


Explore More Data Science and Machine Learning Projects for Practice. Fast-Track Your Career Transition with ProjectPro
Let’s apply K-Means to re-plot this cute puppy
We’ll start K-Means algorithm using our wssplot() function which we created earlier. From the WSS plot below we can see that taking K=5 should suffice our purpose.
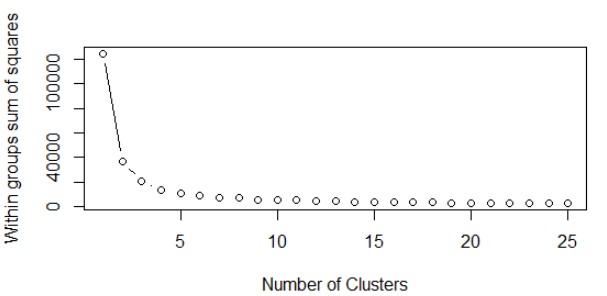
Let’s run the K-Means with ‘5’ clusters
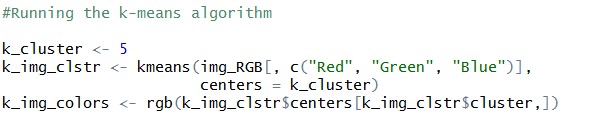
Final step:
Once we get the clusters, next step is to plot the image again with only five colors, which are nothing but centers of the clusters.
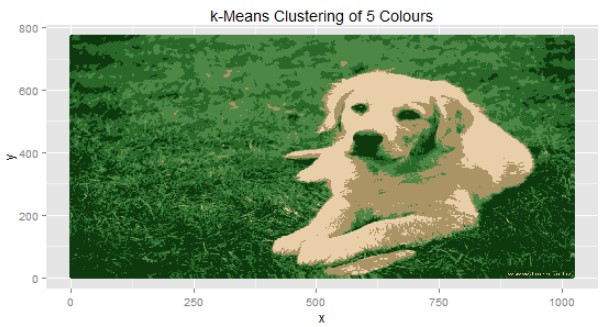
You can see that K-Means has helped us to re-create image using only 5 colors. Can you identify those five colors?
|
R |
G |
B |
Color |
|
0.299311 |
0.530259 |
0.270419 |
|
|
0.915511 |
0.812088 |
0.661382 |
|
|
0.053864 |
0.222914 |
0.056694 |
|
|
0.668194 |
0.58082 |
0.397249 |
|
|
0.16147 |
0.409033 |
0.16128 |
|
Few other concepts
When it comes to clustering there are a few key terminologies which we encounter with. i.e.,
- With-in-Sum-of-Squares (WSS): WSS is the total distance of data points from their respective cluster centroids
- Total-Sum-of-Squares (TSS): TSS is the total distance of data points from global mean of data, for a given dataset this quantity is going to be constant
- Between-Sum-of-Squares (BSS): BSS is the total weighted distance of various cluster centroids to the global mean of data
- R2: R-Square is the total variance explained by the clustering exercise. i.e., BSS/ TSS
We expect our clusters to be tight and homogeneous hence WSS should be lower and BSS should be higher. R2 can be used to assess the progress among different iterations, we should select iteration with maximum R2
Codes
Example 1: With Iris Dataset
#Loading iris dataset
data(iris)
#Viewing the head of iris
head(iris)
#Removing "Species column"
iris_2<-iris[-5]
head(iris_2)
#Standardize data
iris_3<-as.data.frame(scale(iris_2))
head(iris_3)
#Checking Mean and SD of data before and after standardization
sapply(iris_2,mean)
sapply(iris_2,sd)
sapply(iris_3,mean)
sapply(iris_3,sd)
library(NbClust)
# creating function wssplot
wssplot <- function(data, nc=15, seed=1234){
wss <- (nrow(data)-1)*sum(apply(data,2,var))
for (i in 2:nc){
set.seed(seed)
wss[i] <- sum(kmeans(data, centers=i)$withinss)}
plot(1:nc, wss, type="b", xlab="Number of Clusters",
ylab="Within groups sum of squares")}
# calling function wssplot()
wssplot(iris_3,nc=30,seed=1234)
# fitting the clusters
iris_kmeans<-kmeans(iris_3,7)
iris_kmeans$centers
iris_kmeans$size
iris$clstr<-iris_kmeans$cluster
# cross-validation with original species available in data
iris$clstr<-iris_kmeans$cluster
table(iris$Species,iris$clstrExample 2: Image compression
#Installing library
install.packages("jpeg")
#Loading library into R Session
library("jpeg")
#Read image
dog_img<-readJPEG("pet_animal_dog.jpg")
# Obtain the dimension i.e (Pixels, Color Values)
img_Dm <- dim(dog_img)
# Lets assign RGB channels to a data frame
img_RGB <- data.frame(
x_axis = rep(1:img_Dm[2], each = img_Dm[1]),
y_axis = rep(img_Dm[1]:1, img_Dm[2]),
Red = as.vector(dog_img[,,1]),
Green = as.vector(dog_img[,,2]),
Blue = as.vector(dog_img[,,3])
)
library(ggplot2)
# Plot the image
ggplot(data = img_RGB, aes(x = x_axis, y = y_axis)) +
geom_point(colour = rgb(img_RGB[c("Red", "Green", "Blue")])) +
labs(title = "Original Image") +
xlab("x-axis") +
ylab("y-axis")
#Show time - Let's start the clustering
#Running the WSSPLOT function again
wssplot <- function(data, nc=15, seed=1234){
wss <- (nrow(data)-1)*sum(apply(data,2,var))
for (i in 2:nc){
set.seed(seed)
wss[i] <- sum(kmeans(data, centers=i)$withinss)}
plot(1:nc, wss, type="b", xlab="Number of Clusters",
ylab="Within groups sum of squares")}
wssplot(img_RGB[c(3,4,5)],25)
#running the k-means algorithm
k_cluster <- 5
k_img_clstr <- kmeans(img_RGB[, c("Red", "Green", "Blue")],
centers = k_cluster)
k_img_colors <- rgb(k_img_clstr$centers[k_img_clstr$cluster,])
#plotting the compressed image
ggplot(data = img_RGB, aes(x = x_axis, y = y_axis)) +
geom_point(colour = k_img_colors) +
labs(title = paste("k-Means Clustering of", k_cluster, "Colours")) +
xlab("x") +
ylab("y")